인텔 랩, 데이터 장벽 제거해 뇌종양 검출 33%↑
기사입력 2022.12.06 11:11

▲연합 학습(Federated Learning) 개념도 (이미지 - 인텔)
프라이버시 보존 AI, 연합 학습으로 의료 혁신 가속화
6대륙 71개 기관 6,000 GBM 환자 데이터 광범위 학습
6대륙 71개 기관 6,000 GBM 환자 데이터 광범위 학습
의료 분야에서는 미국 의료정보보호법(HIPAA)과 같은 국가별 데이터 개인정보보호법으로 인해 오랫동안 데이터 접근성 이슈가 존재해왔다. 따라서, 환자의 의료 정보를 훼손하지 않으면서 필요한 규모의 의료 연구 및 데이터 공유가 거의 불가능하다는 문제가 있었다.
이에 연합 학습 하드웨어 및 소프트웨어를 통해 개인정보보호 규정을 준수하고 컨피덴셜 컴퓨팅(confidential computing)을 통해 데이터 무결성, 개인정보보호 및 보안성을 유지하는 프라이버시 보존 AI가 개발돼 향후 의료 기술 발전에 활용될 전망이다.
■ 인텔 SGX, 데이터 공유 장벽 제거
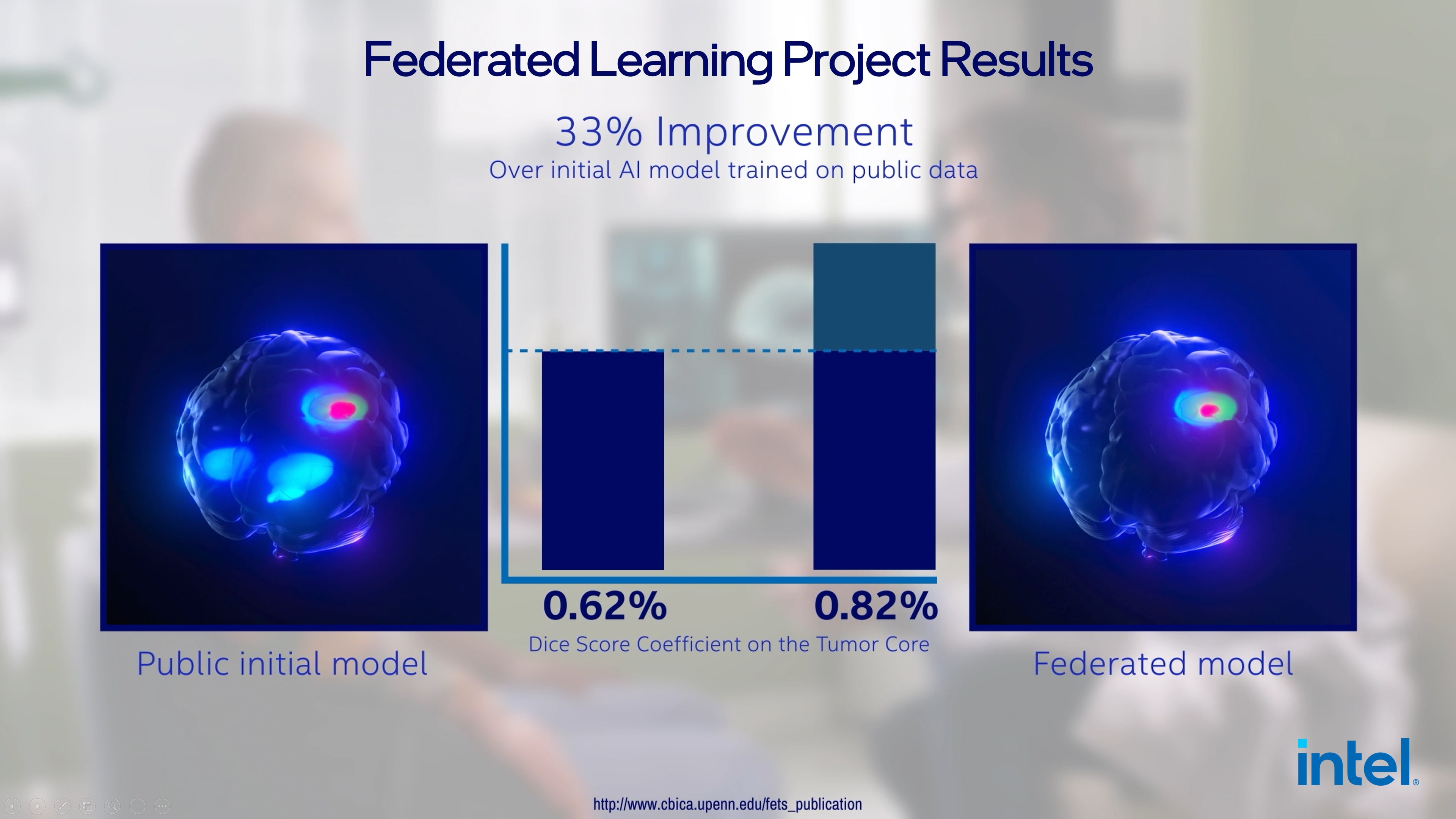
▲연합 학습 기반의 AI 모델은 기존 공개된 데이터로 학습 한 AI 대비 33% 높은 탐지율을 기록했다. (이미지 - 인텔)
본 연구는 6개 대륙에 걸쳐 71개 기관에서 조사한 광범위한 데이터 세트를 통해 진행하는 등 가장 대규모의 의료 분야 연합 학습 부문 연구로, 뇌종양 탐지를 33% 개선할 수 있는 능력을 입증했다.
인텔과 펜 메디슨이 수행한 연구는 인텔 소프트웨어 가드 익스텐션(SGX)과 인텔 연합 학습 기술을 사용한 분산 시스템에서 대량의 데이터를 처리하는 방식으로 진행됐다. 인텔 SGX는 이와 유사한 암 및 질병 연구에서 협업을 제한했던 데이터 공유 관련 장벽을 제거하는 역할을 했다.
분산 시스템은 데이터 소유자의 인프라 내부에 원본 데이터를 보관하며, 해당 데이터를 바탕으로 한 모델 업데이트만 중앙 서버 또는 애그리게이터(aggregator)로 전송함으로써 수많은 데이터 개인정보보호 문제를 해결한다.
제이슨 마틴(Jason Martin) 인텔 랩 수석 엔지니어는 “펜 메디슨과의 연구에서 알 수 있듯이 연합 학습은 다양한 영역, 특히 의료 분야에서 엄청난 잠재력을 가지고 있다”며 “민감한 정보와 데이터를 보호하는 기능은 특히 데이터 세트에 접근할 수 없는 경우, 향후 연구와 협업에 대한 기회를 조성한다”하고 설명했다.
이번 연구 보고서 수석 저자이자 펜 메디슨 병리학·진단검사의학과 방사선학부 조교수인 스피리돈 바카스 박사(Spyridon Bakas PhD)는 “이번 연구를 통해 연합 학습은 패러다임 전환을 통해 다양한 기관에서 방대하고 가장 다양한 교모세포종 환자들의 정보를 이동시키지 않고도 해당 정보에 접근할 수 있게 함으로써 많은 기관들이 협업할 수 있는 잠재력을 보여줬다”며 “머신러닝 모델에 더 많은 데이터를 제공할수록 정확도가 향상되며, 결국 교모세포종과 같은 희귀한 질병을 이해하고 치료하는 역량을 한층 강화할 수 있다”고 말했다.
이러한 인텔과 펜 메디슨의 연구 결과는 동료 심사 학술지인 네이처 커뮤니케이션즈(Nature Communications)에 수록됐다.
■ 6,000 GBM 환자 데이터 학습
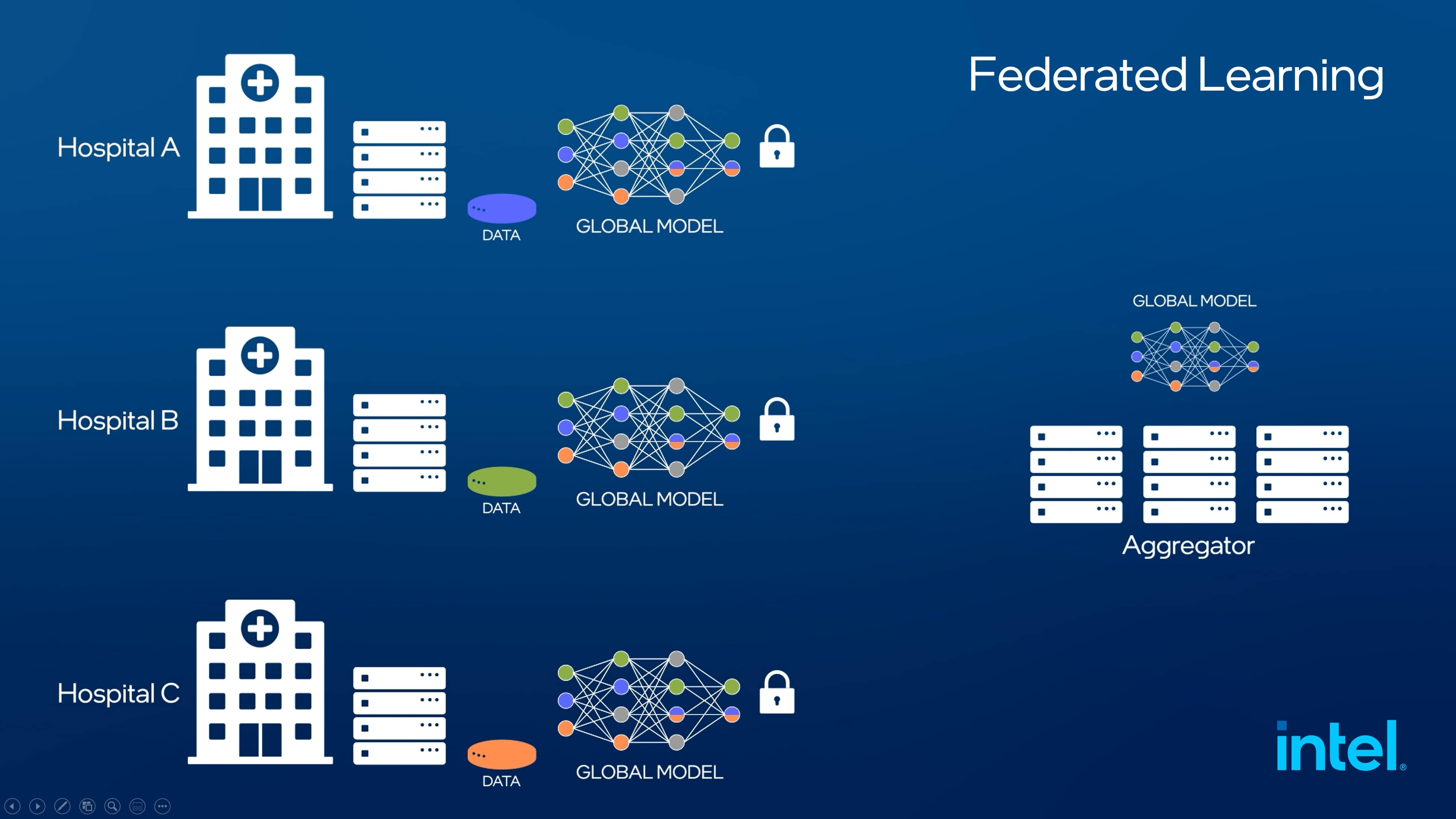
▲연합 학습을 통해 원본 데이터가 저장되어 있는 곳에서 모델을 업데이트 한 뒤 이를 중앙 서버로 이동한다. (이미지 - 인텔)
질병 퇴치를 위한 연구를 발전시키려면 대부분의 경우 하나의 기관에서 생성할 수 있는 임계값을 초과하는 대규모 데이터 세트에 연구진이 접근할 수 있어야 한다.
이번 연구는 대규모 연합 학습의 효과와 다중 데이터 고립 현상이 해소될 때 의료 산업이 실현할 수 있는 잠재적인 이점을 보여준다. 즉, 질병 조기 탐지를 포함해 삶의 질 향상 또는 환자 수명 증가 등의 성과를 달성할 수 있다.
롭 앤덜(Rob Enderle) 앤덜 그룹 수석 분석가는 “세상의 모든 컴퓨터는 충분한 양의 데이터를 분석하지 않고는 많은 성과를 만들어낼 수 없다”며 “이미 확보된 수많은 데이터를 분석할 수 없는 상황때문에 AI가 보장하는 대규모 의료 혁신이 상당히 지연됐다”고 토로했다. 그는 “이번 연합 학습 연구는 AI가 난치병과 싸우는 가장 강력한 도구로 발전해 그 잠재력을 발휘할 수 있는, 실행 가능한 길을 보여주었다”고 말했다.
인텔과 펜 메디슨은 2020년 표준 치료 후 14개월 만에 중위 생존을 보이는 가장 흔하고 치명적인 성인 뇌종양인 교모세포종(GBM)이라는 희귀한 형태의 암의 종양 검출 역량과 치료 결과를 개선하기 위해 연합 학습 연구 및 사용에 협력하기로 발표했다.
지난 20년간 치료 방법이 확대되었지만 전반적인 생존율은 개선되지 않았다. 먼저 펜 메디슨과 71개의 국제 의료 및 연구 기관은 인텔의 연합 학습 하드웨어와 소프트웨어를 사용해 희귀 암의 경계를 탐지하는 방법을 개선했다.
그리고 방사선과 의사들은 종양의 경계를 결정하고 종양 또는 ‘종양핵’의 ‘수술 가능 영역’의 식별을 개선하기 위해 FeTS(Federated Tumor Segmentation)라는 새로운 최첨단 AI 소프트웨어 플랫폼을 사용했다.
방사선과 의사들은 데이터에 주석을 달았고 기계 학습 알고리즘을 학습하기 위한 오픈 소스 프레임워크인 오픈 연합 학습(OpenFL)을 사용하여 연합 학습을 실행했다. 이 플랫폼은 지금까지 가장 큰 뇌종양 데이터 세트인 6개 대륙에 걸쳐 6,000 GBM 환자의 데이터에 대해 학습됐다.
인텔 랩과 펜 메디슨은 이 프로젝트를 통해 연합 학습을 사용해 데이터로부터 지식을 얻기 위한 개념 증명을 만들었다. 이 솔루션은 특히 다른 유형의 암 연구에서 의료나 그 이상에 상당한 영향을 미칠 것으로 예상된다.
특히, 인텔은 OpenFL 오픈소스 프로젝트를 개발해 고객이 실제 교차 사일로(silo) 연합 학습을 채택하고 인텔 SGX에 배포할 수 있도록 지원한다. 또한, 새로운 FeTS 이니셔티브는 지속적인 개발을 위한 플랫폼을 제공하고 깃허브에서 사용할 수 있는 FeTS 플랫폼과 인텔의 OpenFL 오픈 소스 툴킷과의 협업을 장려하기 위해 협업 네트워크를 설립했다.
많이 본 뉴스
[열린보도원칙] 당 매체는 독자와 취재원 등 뉴스이용자의 권리 보장을 위해 반론이나 정정보도, 추후보도를 요청할 수 있는 창구를 열어두고 있음을 알려드립니다.
고충처리인 장은성 070-4699-5321 , news@e4ds.com



.jpg)

.jpg)

